Siglec-7 (also known as p75/AIRM1 or CD328) is a human CD33-related Siglec, described as an inhibitory receptor primarily expressed on natural killer (NK) cells and present on monocytes, dendritic cells, granulocytes and some CD8+ T cells.(Gieseke et al., 2012; Maggi & Moretta, 2020; Shao et al., 2016) It shares high sequence homology with Siglec-9. (Fraschilla & Pillai, 2017) Siglec-7 has three Ig-like extracellular domains (one N-terminal V-set and two C2-set Ig domains) and a cytoplasmic tail containing one ITIM and one ITIM-like domains, involved in the inhibitory signal inside the cell. (Maggi & Moretta, 2020; Yamakawa et al., 2020)
Tumor cells exploit the negative signalling initiated by Siglec-7: overexpressing Siglec-7 ligands, these cells evade the immune system, blocking the NK cell-mediated lysis.(Maggi & Moretta, 2020) For this reason, Siglec-7 has gained attention as a potential target for the treatment of cancer.
Some studies showed that anti-Siglec-7 mAb inhibits the proliferation of chronic myeloid leukemia (CML) cells in vitro and the growth of mast cells in vitro and mice, confirming its potential for cancer treatment and diseases linked to mast cells, such as mastocytosis. (Landolina et al., 2020) Next to immune cells, Siglec-7 occurs on platelets, where it causes their apoptosis. (Nguyen et al., 2014) Siglec-7 occur on β-cells of the pancreas, too: the overexpression on these cells leads to a reduction of β-cells dysfunction. It is downregulated in type 1 and 2 diabetes. (Läubli & Varki, 2020)
Siglec‐7 preferentially binds to α2-8‐linked di-sialylated carbohydrate structures, but it can also bind to α2-6- and α2-3-linked sialic acid-containing glycans. (Gieseke et al., 2012 ; Legrand et al., 2019 ; Movsisyan & Macauley, 2020) Among the six crystal structures of Siglec-7, one has been co-crystallized with the ganglioside GT1b (Figure 12). (Attrill et al., 2006)
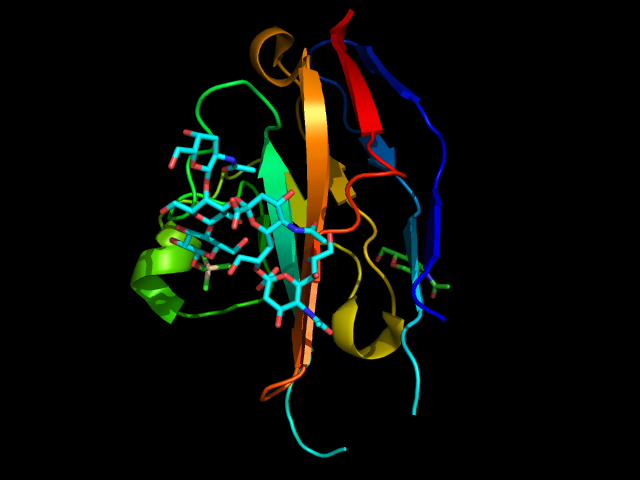
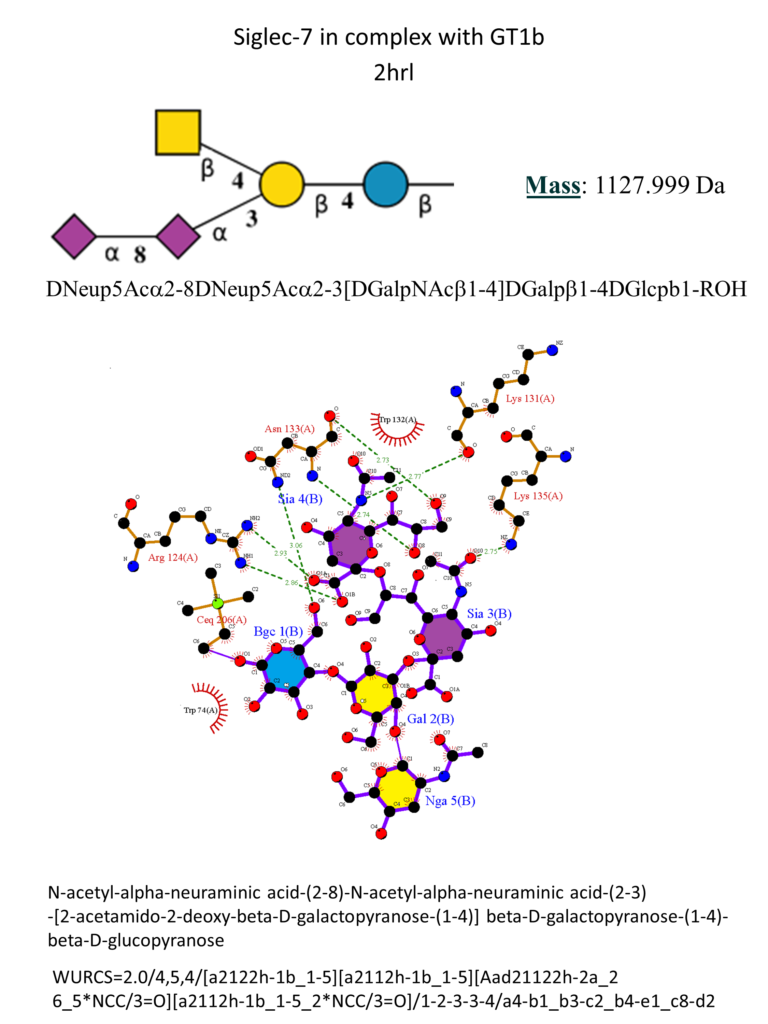
The main interactions involve a salt bridge between the carboxylic acid of the sialic acid and the conserved Arg124, three hydrogen bonds between the hydroxyl groups in positions 5, 8, and 9 and the two amino acids Lys131 and Asn133. Hydrophobic interactions involve glucose and the two amino acids Asn133 and Trp74. (Movsisyan & Macauley, 2020)
Serendipity identified one of the first selective ligands for Siglec-7, but no affinity values are available (Figure 13). Compound 13 presents a xanthene group in position 9 of the sialic acid, which confers selectivity to Siglec-7 by forming π-stacking interactions with the cleft close to position 9. (Rillahan et al., 2013)
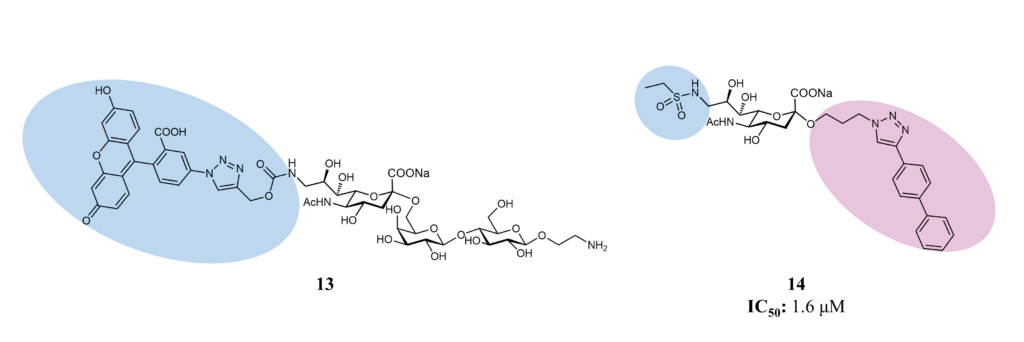
One of the most promising ligands has been developed starting from the methyl-α-Neu5Ac. The introduction of a sulfonamide group in position 9 and an aromatic aglycone in position 2 of the sialic acid increased the affinity of 5000-fold compared to the unmodified reference compound (1.6 μM vs 8000 μM; compound 14, Figure 13). (Prescher et al., 2017)
Further steps towards targeting Siglec-7 have already been made by developing Siglec-7-derived CAR T-cells. They can kill cancer cells by binding to carbohydrates, with a recently found in vitro efficacy. (Lenza et al., 2020)
