The predicted protein models in the AlphaFold protein structure database all lack coordinates for small molecules essential for molecular structure or function: haemoglobin lacks bound heme, zinc-finger motifs lack zinc ions essential for structural integrity, and metalloproteases lack metal ions needed for catalysis. Ligands important for biological function are absent, too; no ADP or ATP is bound to any of the ATPases or kinases.
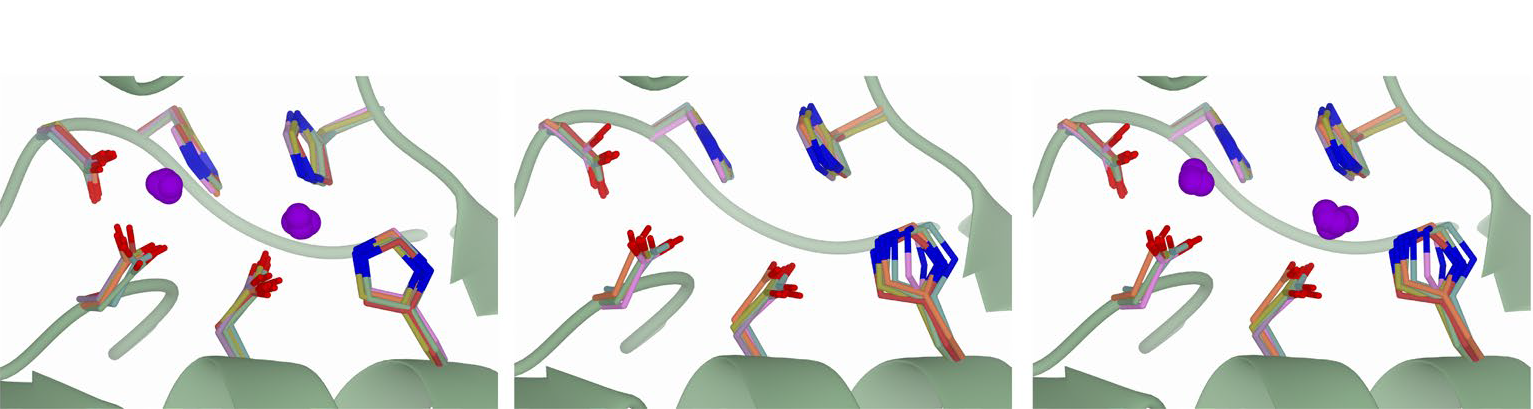
The article presents AlphaFill, an algorithm that uses sequence and structure similarity to ‘transplant’ such ‘missing’ small molecules and ions from experimentally determined structures to predicted protein models. The algorithm was successfully validated against experimental structures. A total of 12,029,789 transplants were performed on 995,411 AlphaFold models and are available with associated validation metrics in the alphafill.eu databank, a resource to help scientists make new hypotheses and design targeted experiments.