Glycans are branched, structurally diverse, and highly flexible biomolecules. These characteristics make glycoanalytics and structural characterization challenging, resulting in often unclear structure-to-function relationships. GlycoShape, currently the largest open- access database of glycan 3D structures from molecular dynamics (MD) simulations, provides an opportunity to fill this information gap. Here, we present GlyContact, an open- source Python package designed and developed to retrieve, process, and analyze glycan 3D structures, from MD, NMR, or X-ray crystallography. We demonstrate that GlyContact can unveil the impact of sequence context on glycan motif structure, (ii) yield a predictive understanding of motif flexibility and surface accessibility on lectin-glycan binding, which improved lectin-binding prediction by ~7%, and (iii) accurately predict torsion angle distribution between disaccharides using von Mises graph neural networks. We envision that GlyContact will allow researchers to explore glycan structures within their 3D space, obtaining insights into their biological functions. GlyContact is available open-access at https://github.com/lthomes/glycontact.
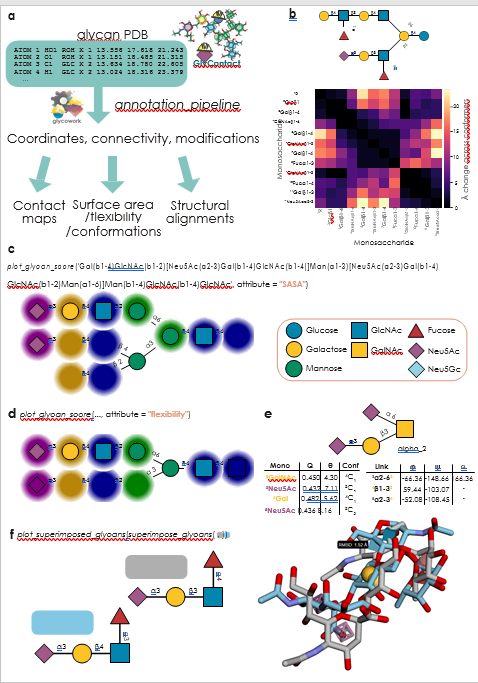
FGlyContact is a powerful platform to process and analyze glycan 3D structures across multiple conformers. a Schematic overview of the main capabilities of GlyContact. Aided by functionality from glycowork, GlyContact extracts glycan-relevant information from PDB files and then can be used to analyze coordinates within and across structures, as well as calculate biophysical parameters, from solvent-accessible surface area (SASA) to monosaccharide conformations, as well as glycan 3D alignments. b Monosaccharide contact map of example glycolipid. Across all beta conformers of the shown glycan, using the draw_contact_map and inter_structure_variability_table functions, we compared how stable the contacts between two monosaccharides were across conformers.