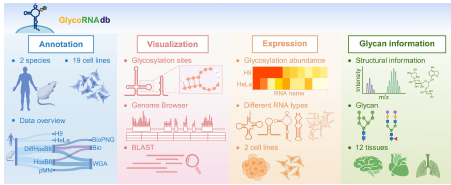
Glycosylated RNAs (glycoRNAs), recently identified as cell-surface molecules modified by complex N-glycans, mark a significant breakthrough in RNA biology, with emerging functions in immune regulation and intercellular communication. To fill the gap of a dedicated resource, the authors developed GlycoRNA db (http://www.glycornadb.com), the first comprehensive database that integrates experimentally supported, specifically enriched glycoRNA sequences with associated glycosylation sites, secondary structures, expression profiles, and glycan composition from human and mouse tissues and cell lines. Data were curated from public datasets and processed through specialized pipelines to identify enriched glycoRNAs. The database contains 3379 curated entries, each annotated with sequence data, structure (experimentally determined or R2DT-predicted), glycosylation sites, expression patterns, and detailed glycan information (composition, linkages, and tissue distribution). Built with a FAIR-compliant architecture, GlycoRNA db offers advanced search options: BLAST-based homology search, JBrowse-based genome browser, and interactive visualizations of structures, modification maps, heatmaps, and glycan chemistry. It enables exploration of cell-type-specific modification patterns, sequence–structure relationships, and modifications near glycosylation sites, such as acp3U, as demonstrated by case studies on snoRNAs and tRNAs. By consolidating scattered glycoRNA data into a centralized, analytically rich platform, GlycoRNA db serves as an essential resource to accelerate research into the biological functions and therapeutic potential of this new class of RNA molecules.