GLYCAM Bacterial Carbohydrate Builder: a web-tool for modelling 3D structures of bacterial glycans
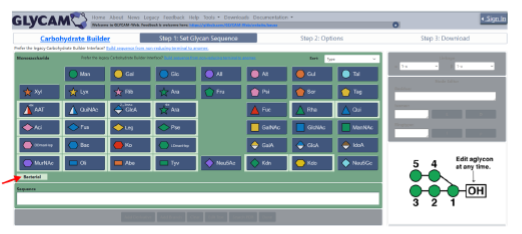
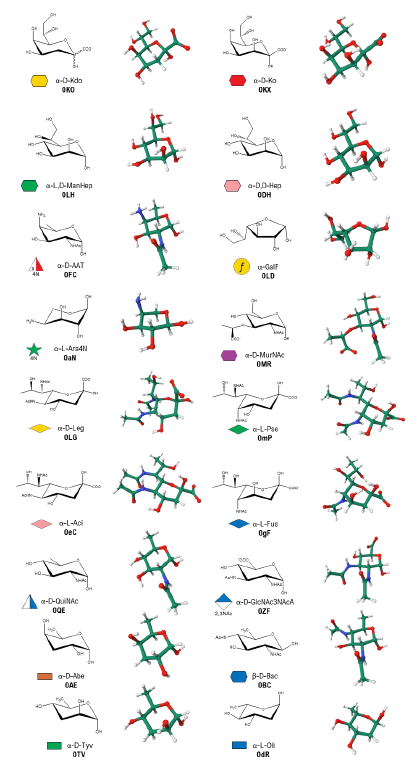
The authors present the GLYCAM Bacterial Carbohydrate Builder (https://glycam.org/cb), an enhanced version of the GLYCAM-Web Carbohydrate Builder1 structure modeler that supports modeling bacterial glycans and enables straightforward generation of three-dimensional structural models. The tool integrates bacterial monosaccharide parametrizations into a curated, user-friendly web-based resource. It provides an intuitive interface for generating carbohydrate sequences and produces 3D structural models in PDB format, along with the input files required to perform molecular dynamics simulations with the AMBER software package. The current implementation includes a library of 18 bacterial monosaccharidesthat can be used in combination with the already-parametrized eukaryotic sugars to construct complex bacterial glycans. Common derivatives, including acetylation, methylation, and sulfation, are also supported. By validating and integrating bacterial sugar parameters into the GLYCAM-Web Carbohydrate Builder, this work reduces the technical barriers associated with bacterial glycan modeling and facilitates computational studies of complex bacterial glycoconjugates.