As outlined above, much of what is known about the mechanism of HA cross-linking comes from studies on the expansion of cumulus matrix. In particular, it was demonstrated that the presence of Hyaluronan(HA), Tumor Necrosis Factor-Stimulated Gene-6 (TSG-6), Inter-α-inhibitor (IαI) and Long Pentraxin 3 (PTX3) is essential for COC matrix expansion and stabilization Zhuo et al., 2001 Salustri et al. 2004Scarchilli et al., 2007
Hyaluronan
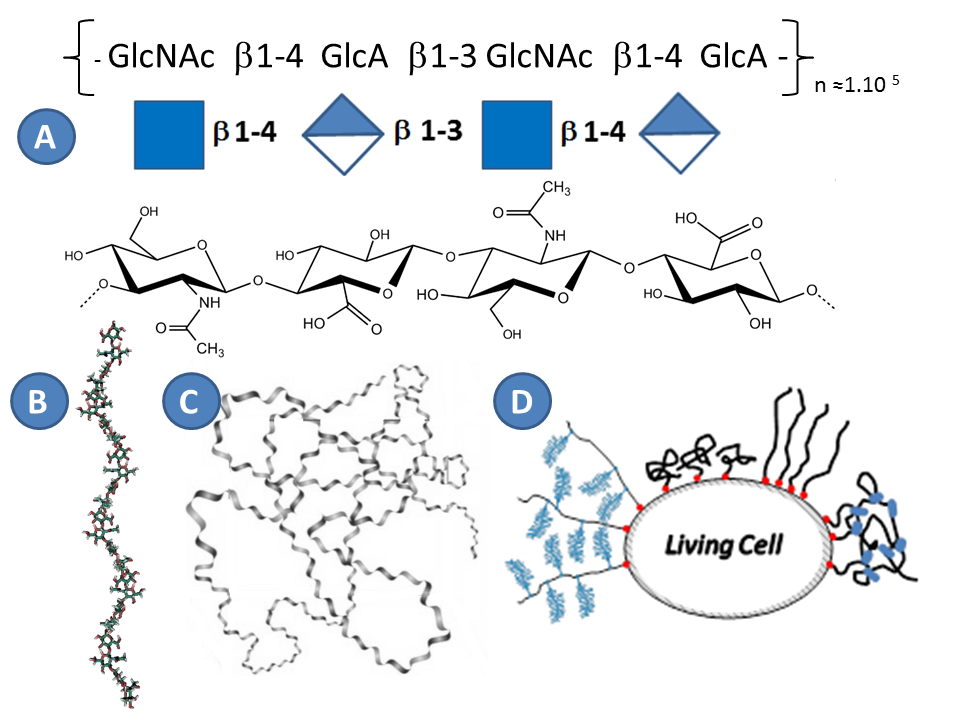
Hyaluronan (HA), or hyaluronic acid, is the main carbohydrate component of the PCM, and defined together with collagen and fibronectin the entire structure of extracellular matrixToole, 2004. HA was discovered in 1934 by Meyer in the vitreous humor of the eye Meyer & Palmer, 1934, and belongs to the family of glycosaminoglycans. In contrast to all other carbohydrates, HA is synthesized directly on the plasma membrane and thus it does not undergo typical chemical modifications in the golgi apparatus as other glycosaminoglycans Weissmann, 1954. HA hence has a simple structure that has been highly preserved during evolution Laurent, 1998. As all glycosaminoglycans, HA is linear. HA is a regular polymer of disaccharides, with the repeating unit consisting of glucuronic acid and N acetylglucosamine (Figure 4). The number of disaccharides, n, per chain can reach 10000 Laurent, 1998. With each disaccharide (HAds) having a molecular weight of 378 Da, the polymer chain can hence reach a molecular weight of 4MDa. The length of HAds is approximately 1 nm, and thus a single HA chain can reach a length of 10 µm if it is fully extended (i.e. the so-called contour length) Itano et al., 2002 Tammi et al., 2002.
From a polymer physics point of view, HA is well described as polyelectrolyte Albersdörfer & Sackmann, 1999 Cleland, 1984 Attili, 2012. The pKa of the glucuronic acid is about 3-4, and thus at physiological pH the carboxyl groups are predominantly ionized, rendering the HA chain highly negatively charged Hascall, 1997. Intramolecular hydrogen bonds stiffen the chain Sheehan & Atkins, 1983. Because the HA chains are relatively stiff and repel each other, and because of thermal energy, isolated HA chain in solution adopts the conformation of a swollen random coil. For the typical size of HA polymers, the water content in such a coil is around 99.9% Cleland, 1984 Cleland & Wang, 1970.
A single HA chain is very long and hence can have hundreds of binding sites for hyaladherins Attili et al., 2012. Hyaladherins typically bind to between 6 and 20 monosaccharides of HADay & Prestwich, 2002. The persistence length of an HA chain, a measure of chain flexibility, is about 7 nm Attili et al., 2012Takahashi et al., 1999. On the length scale of a hyaladherin binding cite, the hyaluronan chain therefore appears rather rigid
The concentration of HA in tissues varies from 0.01 µg/ml in blood plasma to about 4 mg/ml in synovial fluid and umbilical cord Fraser et al., 1997. In the expanded COC matrix, for example, the HA concentration is about 0.5 mg/ml Salustri et al., 1998. At this concentration, the swollen random coils formed by individual HA chains start to overlap and hence pervade the entire solution, forming a transient meshwork of entangled polymer chains. The typical size of a transient mesh in such an meshwork would be about 50 to 100 nm. As discussed above, the HA meshwork in the COC matrix is stabilized by protein cross-linkers. The mesh size in a cross-linked meshwork should depend on the density and stability of cross-links.
Although HA has a several micrometers long contour length and can adopt a random coil conformation in solution occupying a large volume (the radius of gyration, a measure for the coil size, for longest HA chains is about 500 nm), this alone cannot explain the origin of the extended matrices with thicknesses of several micrometers. In the literature different models were proposed to explain the formation of the thick coats (Figure 4D) :
- 1) the PCM might consist of multiple layers of cross-linked and entangled HA chains Cohen et al., 2006
- 2) the cell membrane might be extremely rough to form microvilli Kulti et al., 2006Rilla et al., 2008
- 3) it is also possible that there is a formation of a dense HA brush, because on the cell surface HA chain remains attached via HA synthases Knudson, 1993 Evanko et al., 1999.
In addition, steric repulsion between bottle-brush shaped proteolgycans, like versican of the COC matrix Russel et al., 2003can also affect HA configuration by stretching individual HA chains in similar manner as aggrecan Knudson, 1993 Lee et al.,1993 Attili & Richter, unpublished dat. The mentioned mechanisms may be specific to a given tissue or they might act together. Overall, the interactions between HA and hyaladherins, but not HA alone, dictate the final properties of the PCM such as thickness, permeability and mechanical properties.
Tumor Necrosis Factor-Stimulated Gene-6 : TSG-6
The secreted product of tumor necrosis factor-stimulated gene-6 (TSG-6) is a 35 kDa protein composed of a Link and a CUB module. The mRNA encoding the protein was originally identified following tumor necrosis factor treatment of human fibroblasts Lee et al., 1992Maier et al., 1996. The protein TSG-6 has been reported in the synovial fluid of arthritis patients at concentrations of nM or 0.6 μM Wisniewski et al., 1993, but it is not constitutively expressed under healthy conditions. The precise role of TSG-6 in inflammation is still unclear, though it was demonstrated that it protects from the cartilage destruction in the mice model of arthritis Glant et al., 2002Mindrescu et al., 2002Szanto et al., 2004. In ovulation, TSG-6 was shown to be critical for COC matrix expansion Fulop et al., 2003.
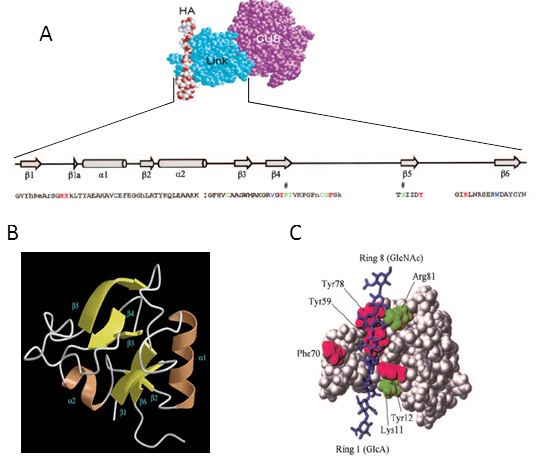
The Link module contains the hyaluronan binding domain which is conserved among HA binding proteins Blundell et al., 2003Blundell et al., 2005. The tree dimensional structure of a protein construct containing essentially the Link module (Link_TSG-6) has been determined in HA-free and HA-bound conformations Khoda et al., 1996 (Figure 5). Link_TSG-6 has a molecular weight of 10.9 kDa and dimension of 3 × 5 × 2 nm3 (Protein Data Bank code 2WNO).
It is demonstrated that HA8 is the minimal sugar length that can stably bind to the protein. By site directed mutagenesis five main amino acids, mostly aromatic residues (i.e. Lys11, Tyr12, Tyr59, Phe70 and Tyr78), were identified to be involved in HA binding Khoda et al., 1996. In addition, Arg81 and Lys81 were proposed to make a salt bridge with GlcA on the basis of NMR data Blundell et al., 2005. The presence of salt bridges was confirmed by isothermal calorimetry performed over a range of NaCl concentrations Blundell et al., 2002.
The CUB module is not known to have HA binding properties. Its structure is currently not solved, but molecular modeling suggested that it is involved in the bonding of TSG-6 to IαI Carrette et al., 2001. Also, it was demonstrated that the CUB domain is responsible for the interaction between TSG-6 and fibronectinKuznetsova et al., 2008.
Small angle X-ray scattering data suggested full length TSG-6 to form dimers in the presence of HA octasaccharides in solution Day et al., 2004. The TSG-6 dimer was estimated to be 13±1 nm in length. Based on this finding, self-association between TSG-6 monomers was proposed as one of the mechanisms to cross-link HA matrices and thereby to stabilize the matrix against destruction like in arthritis Day & de la Motte, 2005(Figure 2 A).
Inter-α-inhibitor ( IαI)
Inter-α-inhibitor (IαI) is a proteoglycan. It is composed of two heavy chains (HC1 and HC2) and the serine protease inhibitor bikunin Enghild et al., 1989Salier et al., 1996. The HCs are interconnected with the bikunin subunit by a chondroitin sulfate (CS) chain. The CS chain is 12 to 18 disaccharides long. The HC1 and HC2 subunits ( 80 kDa each) are located in close proximity to each other, in a region of less sulfated CS, with HC1 attached farther away from bikunin than HC2 Enghild et al., 1999. The attachment of bikunin to CS is through a common O-glycosidic bond on Ser, while the HCs are attached via ester bonds formed between the carboxyl group of asparatic acid on the HCs and the C-6 carboxyl group of N-acetylgalactosamine in CS Sanggaard et al., 2005. By electron microscopy, it was shown that bikunin has a spherical shape with a diameter of 2 nm, whereas the two HCs have a globular shape (diameter of 11 nm)Blom et al., 1999. IαI is present in blood plasma (the circulating level is 150 to 500 µg/ml Steinbuch, 1976,Potempa et al., 1989, and in extracellular matrix Zhuo et al., 2001Blom et al., 1995Bost et al., 1998.
Its name reflects that IαI was initially extensively studied as a protease inhibitor, but its inhibitory activity on proteases, realized through the bikunin subunit, is rather weak Potempa et al., 1989. It was suggested that IαI functions as a shuttle that traps proteases and then transfers them to the physiological inhibitors Pratt & Pizzo, 1986. An inhibitory activity of bikunin towards hyluaronidases has also been described Pratt & Pizzo, 1986. Most of circulating bikunin though corresponds to IαI, rather than bikunin-CS, presumably due to the longer half-life of intact IαI Zhuo et al., 2001Sugiki et al., 1989.
In the ECM, IαI was found to be associated with HA although it does not bind directly to HA Sugiki et al., 1990Hamemam & Sandson, 1963. In particular, HA can be covalently modified with HCs, and increased modification of HA with HCs of IαI was found in synovial fluid of patients with rheumatoid arthritis compared to healthy patients. The extracted HA containing covalent HC modifications had properties of a gel, suggesting that HCs can cross-link HASuguki et al., 1989. HA•HC complexes have also been found in follicular fluidJessen et al., 1994. Structural analysis of the complexes revealed that HCs are bound to HA via an ester bond between the Asp residue at the C-terminus of HCs and the C-6 hydroxyl group of N-acetylglucosamine in HA, analogous to the original bond between HCs and CS in intact IαIZhao et al., 1995. Because HA in the ECM has rather large contour length, a single chain can accommodate several HCs. The HA length in rheumatoid arthritis synovial fluid is rather heterogeneous, but HA chains of 2 MDa were found to carry 3 to 5 HCs on averageYingsung et al., 2003.
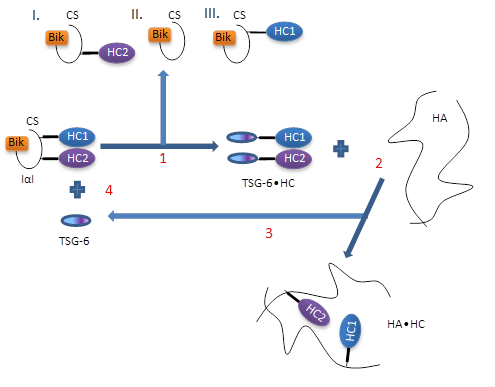
Several requirements need to be fulfilled to allow for covalent HA•HC complexes to form. The first requirement is the presence of an enzyme which facilitates the transesterification reaction. TSG 6 has been identified to fulfill this function Wisniewski & Burgess, 1994, by formation of an intermediate 120 kDa complex with HCs of IαI Rugg et al., 2005. The covalent TSG 6•HC complex has been detected in arthritis synovial fluid, in air pouch exudates and in the COC matrix Wisniewski & Burgess, 1994Mukhopadhya et al., 2004. Rugg et al. Rugg et al., 2005suggested the following mechanism of transfer :
- TSG-6 displaces HCs from IαI, with formation of a TSG-6•HC complex, subsequently ;
- HCs from the TSG-6•HC complex become transferred to HA ;
- TSG-6 is released and engaged into a new interaction with IαI, i.e. it is recycled (Figure 6).
Another important requirement for the formation of HA•HC complexes is the presence of divalent cations, particularly Mn2+ or Mg2+. Both TSG-6•HC complex formation and HCs transfer on HA are metal ion dependent processes Rugg et al., 2005.
- Binding of IαI (or its subunits) to HA is of great impotence for the normal expansion of the COC. In the absence of serum as an IαI source, cultured cumulus cells are not able to organize a stable matrix Chen et al., 1994. While bikunin can be detected in follicular fluid at the point of ovulation, indicating the presence of IαI Jessen & Odum, 2003, COC matrices that were isolated after 24 h contained HCs but not bikunin. This correlates with the proposal that bikunin is released upon formation of HC•HA complexes and does not play any structural role in the HA matrix Jessen et al., 1994Wisniewski et al., 1994Mukhopadhyay et al., 2004. Bikunin knockout mice, even though they were producing free HCs, were not able to form HC•HA complexes or to assemble a functional matrix Zhuo et al., 2001. This highlights that the presence of the intact form of IαI is essential for the formation of HC•HA complexes. In particular, it is important to have the ester bond between CS and HCs which later can be transferred to HA.
Self-association between HCs was hypothesized Day & de la Motte, 2005 based on the findings that HA from synovial fluid of arthritis patients aggregates and gelates at low pH Yingsung et al., 2003. However, for the organization of functional COC matrix, the presence of the three components HA, TSG 6 and IαI is still not sufficientSalustri et al., 2004Scaechilli et al., 2007.
Long pentraxin 3 : PTX3
Long pentraxin 3 (PTX3) is an acute phase glycoprotein produced at sites of inflammation by both somatic and immune cells Breviario et al., 1992Lee et al., 1993Bottazzi et al., 1997. Similar to TSG-6, PTX3 expression can be induced by treatment with interleukin-1 and TNF 102. PTX3 plays an important role in innate immunity, vascular biology and fertility Varani et al., 2002Bottazzi et al., 1997Bottazzi et al., 2009Garlanda et al., 2005. PTX3 forms octamers with a molecular weight of 340 kDa in which the quaternary structure is held together by disulfide bonding. Each protomer of 42.5 kDa contains N- and C-terminal globular domains Bottazzi et al., 1997Inforzato et al., 2008. The N-terminal region (178 amino acids) has a unique structure that has not been found in any other protein. It contains three α-helices which form a coiled-coil assembly Presta et al., 2007. The C-terminal region (203 amino acids) corresponds to a pentraxin-like domain, which is homologous to the short pentraxins C-reactive protein and serum amyloid P component Bottazzi et al., 1997Inforzato et al., 2008Goodman et al., 1996Inforzato et al., 2006.
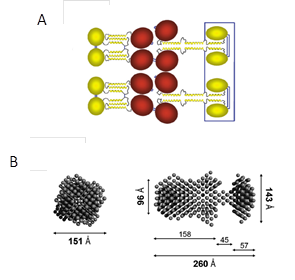
Structural analysis of recombinant PTX3 revealed that it is an elongated, asymmetric molecule with two lobes of tetramers connected via a stalk part (Figure 7)Inforzato et al., 2010. The Cys residues responsible for the quaternary organization of PTX3 are located within the N-terminal domainInforzato et al., 2010. The structural complexity of PTX3 explains the existence of the broad spectrum of binding ligands among which are TSG-6 Salustri et al., 2004, Leali et al., 2012 and HCs of IαI Scarchilli et al., 2007Ievoli et al., 2011.
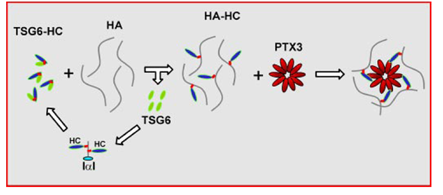
The oligomeric status of PTX3 is of particular importance for the stabilization of the COC matrix. In the COC matrix, PTX3 was found to be co-localized with HCs of IαI, suggesting that the HA•HC is the binding ligand for PTX3 Scarchilli et al., 2007 (Figure 8). It was demonstrated that Cys/Ser mutated forms of the N-terminal domains of PTX3 that form dimers were not able to rescue COC matrix assembly in PTX3 -/- mice, while the native form which forms tetramers was active in restoring matrix expansion Ievoli et al., 2011. Although the N-PTX3 dimer binds both TSG-6 and IαI, the exact amount of binding site for TSG-6 and IαI on the PTX3 octamer remains unknown. The importance of PTX3 oligomerization for the stabilization of COC matrix suggests PTX3 as a nodal molecule in the cross-linking of the HA matrix.
