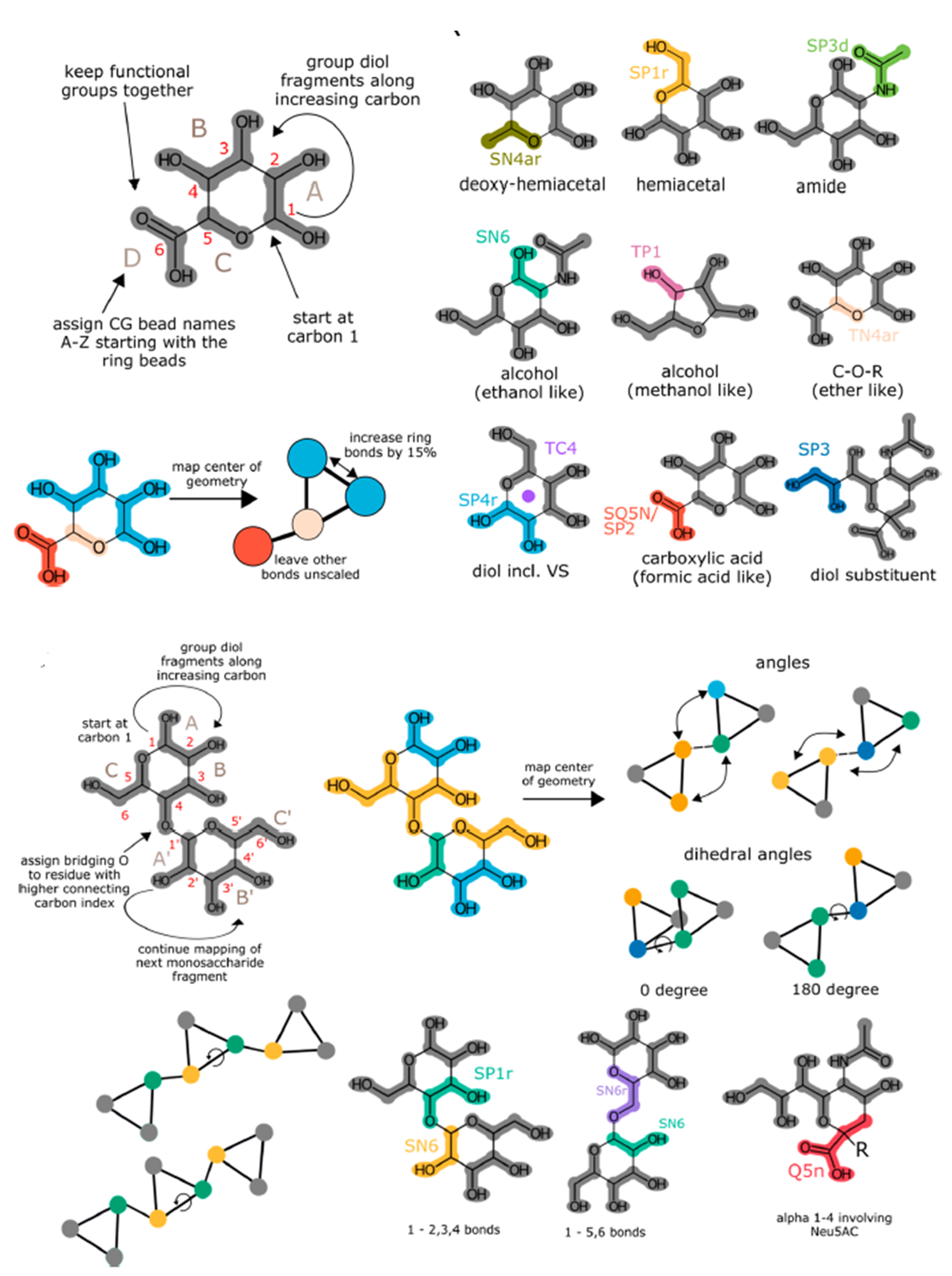
The Martini 3 force field is a full reparametrization of the Martini coarse-grained model for biomolecular simulations. The improved interaction balance allows a more accurate description of condensed phase systems. In the present work, we develop a consistent strategy to parametrize carbohydrate molecules accurately within Martini 3. In particular, we create a canonical mapping scheme which decomposes arbitrarily large carbohydrates into a limited number of fragments. Bead types for these fragments have been assigned by matching the physicochemical properties of mono- and disaccharides. In addition, guidelines for assigning bonds, angles, and dihedrals were developed. These guidelines enable a more accurate description of carbohydrate conformations than in the Martini 2 force field.
The obtained models can accurately reproduce the osmotic pressures of carbohydrate water solutions. The model can correctly differentiate the solubility of the polysaccharide dextran (water-soluble) and cellulose (water-insoluble but soluble in ionic liquids). Finally, we demonstrate that the new building blocks can be applied to glycolipids. The membrane properties and the induced binding of peripheral membrane proteins can also be reproduced. These test cases demonstrate the validity and transferability of the complete parametrization.