Heparin, a naturally occurring glycosaminoglycan, has been found to have antiviral activity against SARS-CoV-. The authors investigated the binding of heparin to the SARS-CoV-2 spike glycoprotein by means of sliding window docking, molecular dynamics simulations, and biochemical assays. The simulations show that heparin binds at long, positively-charged patches on the spike glycoprotein, thereby masking basic residues of both the receptor-binding domain (RBD) and the multifunctional S1/S2 site. Biochemical experiments corroborated the simulation results, showing that heparin inhibits the furin-mediated cleavage of the spike by binding to the S1/S2 site. The simulations indicated that heparin can act on the hinge region responsible for the motion of the RBD between the inactive closed and active open conformations of the spike glycoprotein.
In the simulations of the closed spike homotrimer, heparin binds the RBD and the N-terminal domain of two adjacent spike subunits and hinders opening. From the simulations of open spike conformations, it appears that heparin induces stabilization of the hinge region and a change in RBD motion. Taken together, these results indicate that heparin can inhibit SARS-CoV-2 infection by three mechanisms:
- 1. by allosterically hindering binding to the host cell receptor,
- 2. by direct competition with binding to host heparan sulfate proteoglycan co-receptors,
- 3. by preventing spike cleavage by furin.
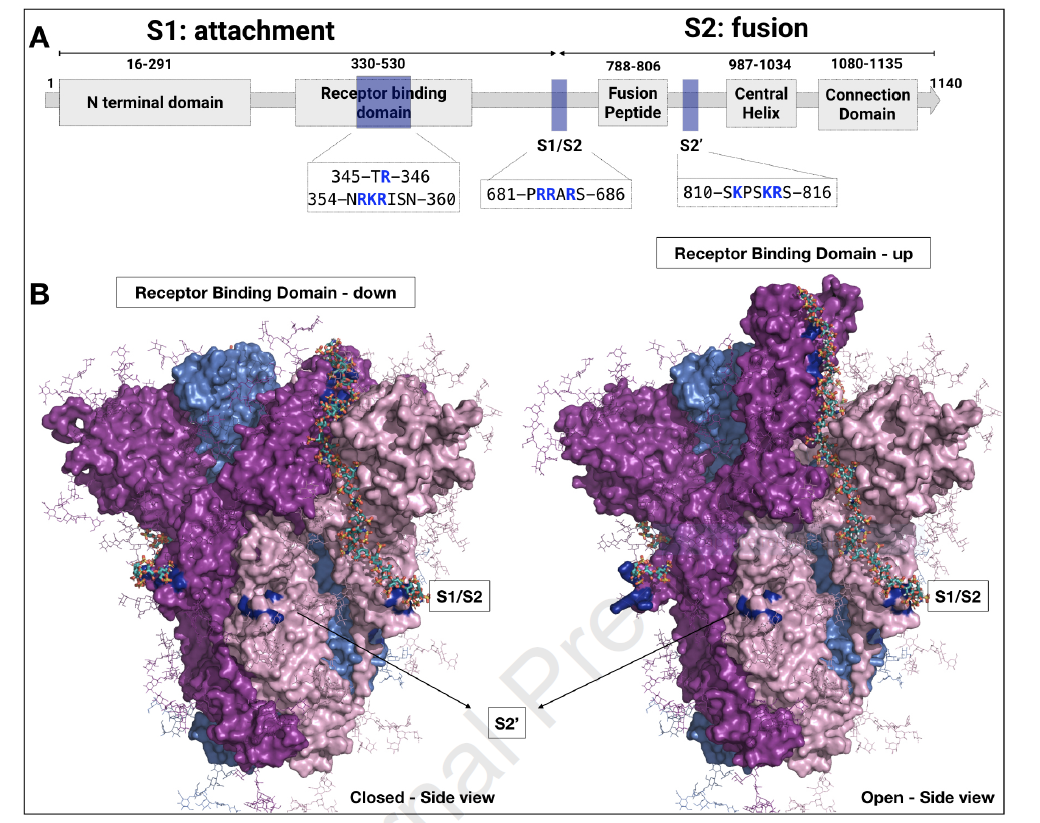
These simulations provide insights into how host heparan sulfate proteoglycans can facilitate viral infection. These results provide a set of molecular bases that could help the rational optimization of heparin derivatives for SARS-CoV-2 antiviral therapy.