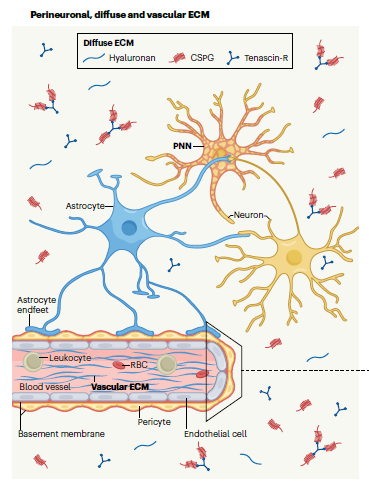
Lipopolysaccharides (LPSs) play a vital role in the elicitation of an innate immune response in all eukaryotes and are recognized as Microbe Associated Molecular Patterns (MAMPs) by the host immune system. LPSs comprise three chemically and genetically distinct regions: the endotoxic moiety termed Lipid A, the core region and
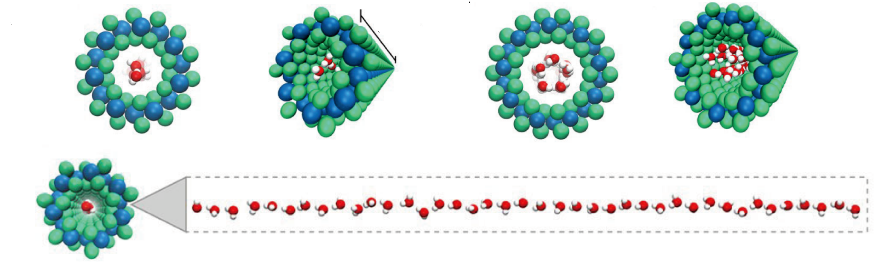
the O-specific polysaccharide (OPS) or O-antigen. In the course of a symbiotic interaction, the bacterial O-antigens play a crucial role in the suppression of the plant’s innate immunity by mimicking host surface carbohydrate antigens. They can also circumvent recognition by building up a chemically unique and tortuous helical supramolecular structure. On the contrary, when the bacterium establishes pathogenic interactions, the OPS is able to elicit plant defense responses.
The research undertaken in the article meant at evaluating the eventual role of the coat of one Lipopolysaccharide (LPS) O-antigen structure of the plant pathogen Rhizobium radiobacter strain TT9 during the infection process on the plant model Arabidopsis thaliana, and to seek a possible relation between the activity measured and the structure of the LPS and/or of the OPS.
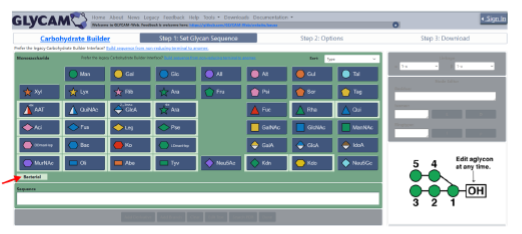
The analyses disclosed the presence of two O-antigens, named Poly1 and Poly2. The repetitive unit of Poly2 defenses by 4-alpha-L-rhamnose linked to a 3-alpha-D-fucose residue. Surprisingly, Poly1 turned out to be a novel type of biopolymer in which the repeating unit is formed by a monosaccharide and an amino acid derivative,. Therefore, the polymer has alternating glycosidic and amidic bonds joining the two units: 4-amino-4-deoxy-3-O-methyl-D-fucose and (2’R,3’R,4’S)-N-methyl-3′,4′-dihydroxy-3′-methyl-5′-oxoproline).

Differently from the O-antigens of LPSs from other pathogenic Gram-negative bacteria, these two O-antigens do not activate the oxidative burst, an early innate immune response in the model plant Arabidopsis thaliana, explaining at least in part the ability of this R. radiobacter strain to avoid host defences during a plant infection process. Pathogenic R. radiobacterT9 develops a unique strategy to carry on the infection process in the plant, with the advantage to prevail over other potential pathogens. An LPS coating not recognised by the host receptor(s) is of great benefit to this pathogen, as its virulence will increase without the induction of the MAMP triggered immunity. The results described in the article bring a new concept as regards to the role of polymer architecture in plant recognition of pathogenic microbes